|
5/16/2023 0 Comments Macvector align two sequencesYou can learn more about sequence alignments on the UniProt help page. You can also run Alignment from within the Basket. All relevant results pages (such as UniProtKB, UniRef, UniParc and tool results) provide an ‘Align’ button to run alignments directly by selecting entries with checkboxes. When youre finished, your page should look something. With the text boxes still selected, group them. Select Align Right and Distribute Vertically. Integrated sequence analysis for the Macintosh Methods Mol Biol. 
Click the Align command, and make sure the Align Selected Objects option is selected. The following kinds of UniProt identifiers are supported: P00750Įach UniProtKB entry which contains both a sequence and one or more isoforms of that sequence, enables you to align the canonical sequence and its isoforms. MUltiple Sequence Comparison by Log-Expectation ( MUSCLE) is computer software for multiple sequence alignment of protein and nucleotide sequences. Hold down the Shift key, then select the text boxes containing Cleaning, Maintenance, Repair, and Restoration. Note – advanced users are given the option of varying the alignment parameters from those given as default. Enter either protein sequences in FASTA format or UniProt identifiers (as above) into the form field.A new empty sequence appears for entering data by typing it manually ( see Note 2 ). To enter a new sequence, select New Sequence from the Sequences menu. 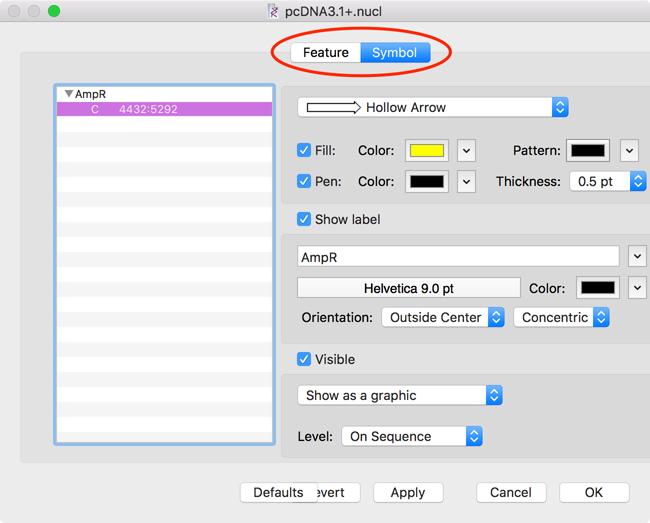
When starting a new layout, only two options will be available under the Sequences menu.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
